What did the scientists discover?
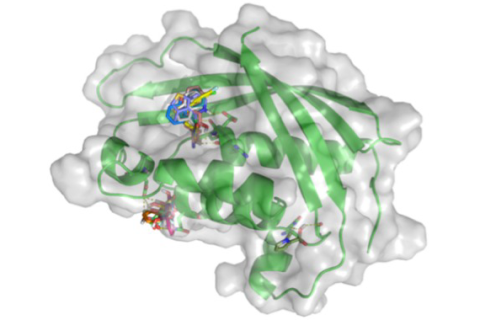
Casu and co-workers used a fluorescence method to screen a library of small molecules ('fragments') to find those which bind to a protein, TraE, essential to the transfer of a piece of genetic material called “plasmid pKM101” from one bacterial cell to another. The figure shows the predicted binding sites for a number of these fragments. But, how do they really bind? To find out, the scientists used X-ray crystallography at CHESS to determine the structures of complexes of TraE with two of the fragments.. The results showed that there are two binding sites for the fragment called 1E6, one of which was unexpected. This novel site presents a new target for designing inhibitors of pKM101.
Impact:
The transfer of plasmids through the process of bacterial conjugation is an important contributor to the increasing frequency of antimicrobial resistance, which makes diseases harder to treat. The researchers have implemented a structure-based method to discover new inhibitors of bacterial conjugation. Although developed for one particular type of bacterial pathogens, the method uses commonly available laboratory instrumentation and can be applied to a wide variety of virulence factors. It has potential to contribute to the development of drugs that reduce the transfer of antimicrobial resistance from one pathogen to another.
Collaborators:
Christian Baron, Université de Montréal, christian.baron@umontreal.ca
Tarun Arya, Université de Montréal
Bastien Casu, Université de Montréal
Benoit Bessette, Université de Montréal
Publication:
“Fragment-based screening identifies novel targets for inhibitors of conjugative transfer of antimicrobial resistance by plasmid pKM101”
By: Bastien Casu, Tarun Arya, Benoit Bessette, and Christian Baron
Publication: Scientific Reports
Vol. 7
Article: 14907
Published on-line: Nov. 2, 2017
Funding:
Canadian Institutes of Health Research (CIHR MOP-84239)
CHESS and MacCHESS grants