What did the Scientists Discover?
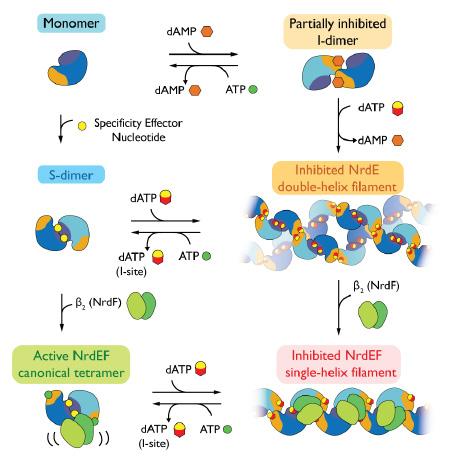
The protein ribonucleotide reductase (RNR) is responsible for synthesis of the building blocks of DNA, making it a very important enzyme in all life. RNRs are often tightly regulated, but the mechanism of the regulation can take different forms in different organisms. In this study, researchers employed small-angle X-ray scattering (SAXS), X-ray crystallography, and cryo-electron microscopy (cryo-EM) to characterize an unexpected mechanism of activation and inactivation that evolved in a class of RNRs found exclusively in bacteria, including many pathogens.
Broader Impacts of this work?
As antibiotic resistance rises, the search for new antibiotic strategies has become imperative. By understanding how an essential enzyme is inactivated in an organism-specific manner, the researchers hope to contribute to the development of new anti-pathogenetic therapies. The research also sheds light on the evolutionary pathways through which this type of regulation may evolve.
Why is this important?
This work mapped out the complicated regulation of an integral enzyme using SAXS, then built upon these results with high-resolution techniques to elucidate the structural changes that determine RNR activation and deactivation. The characterization included the discovery of a remarkable, inhibited enzyme filament. This work can also be a model workflow for future studies of similarly complex and important enzymes both in the RNR family and elsewhere.
Why did this research need CHESS?
The X-ray experiments in this study were done exclusively at CHESS.
At the G1 beamline, the ability to switch between a temperature-controlled in-line FPLC and flow cell allowed the researchers to comprehensively map the many states of the enzyme by SAXS. X-ray diffraction data were collected at F1. Unexpectedly, the functional C-terminal tail was found trapped in the active site via disulfide bonds, allowing its visualization for the first time. The researchers then used X-ray photoreduction to determined a structure of a catalytically important, reduced state.
Collaborators:
Department of Chemistry and Chemical Biology, Cornell University
- William C. Thomas, Audrey A. Burnim, John-Paul Bacik, Nozomi Ando
Department of Chemistry, Princeton University, Princeton
- William C. Thomas, F. Phil Brooks III, Audrey A. Burnim, John-Paul Bacik, Nozomi Ando
Department of Chemistry, Massachusetts Institute of Technology
- JoAnne Stubbe
Institute for Quantitative Biomedicine, Rutgers University
- Jason T. Kaelber
Department of Biochemistry and Molecular Biology, Oregon Health & Science University
- James Z. Chen
Publication Citation:
W.C. Thomas, F.P. Brooks III, A.A. Burnim, J.P. Bacik, J. Stubbe, J.T. Kaelber, J.Z. Chen, N. Ando. Convergent Allostery in Ribonucleotide Reductase; Nature Commun. 10, 2653 (2019) https://doi.org/10.1038/s41467-019-10568-4
How was the work funded?
The research at CHESS is supported under NSF award DMR-1332208 and NIH award GM-103485. This work was supported by NIH grants GM081393 (J.S.) and GM124847 (N.A.) and startup funds from Oregon Health & Science University (J.Z.C.), Princeton University (N.A.), and Cornell University (N.A.).