Since September 2019, the National Science Foundation (NSF) supports the operation of the Center for High Energy X-ray Sciences (CHEXS) at the Cornell High Energy Synchrotron Source (CHESS) as a national user facility. CHEXS consists of a suite of high-energy X-ray beamlines that serve the national user community in specifically targeted research areas in materials research, biology and engineering. Recently, first research articles that were supported by NSF through CHEXS have been published:
Spontaneous Gyrotropic Electronic Order
 Chirality – or handedness – is a symmetry property in nature that can be found in many forms and on many length scales, from the spiral-staircase design of DNA to extended spiral galaxies. Now, a team of researchers has demonstrated spontaneous chiral symmetry breaking by electrons emerging from a parent “charge-density-wave” (CDW) ordered state.
Chirality – or handedness – is a symmetry property in nature that can be found in many forms and on many length scales, from the spiral-staircase design of DNA to extended spiral galaxies. Now, a team of researchers has demonstrated spontaneous chiral symmetry breaking by electrons emerging from a parent “charge-density-wave” (CDW) ordered state.
Nature 578, 545 (2020); DOI: 10.1038/s41586-020-2011-8
CHESS Highlight
Unsupervised learning of dislocation motion
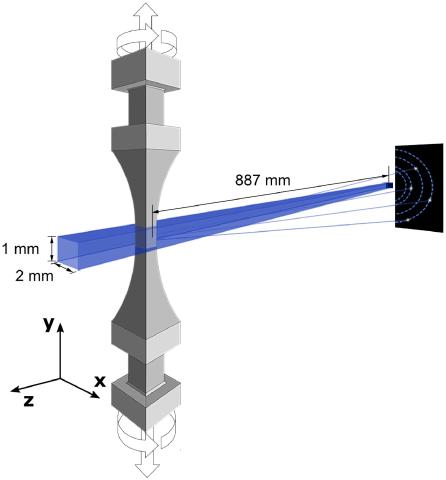
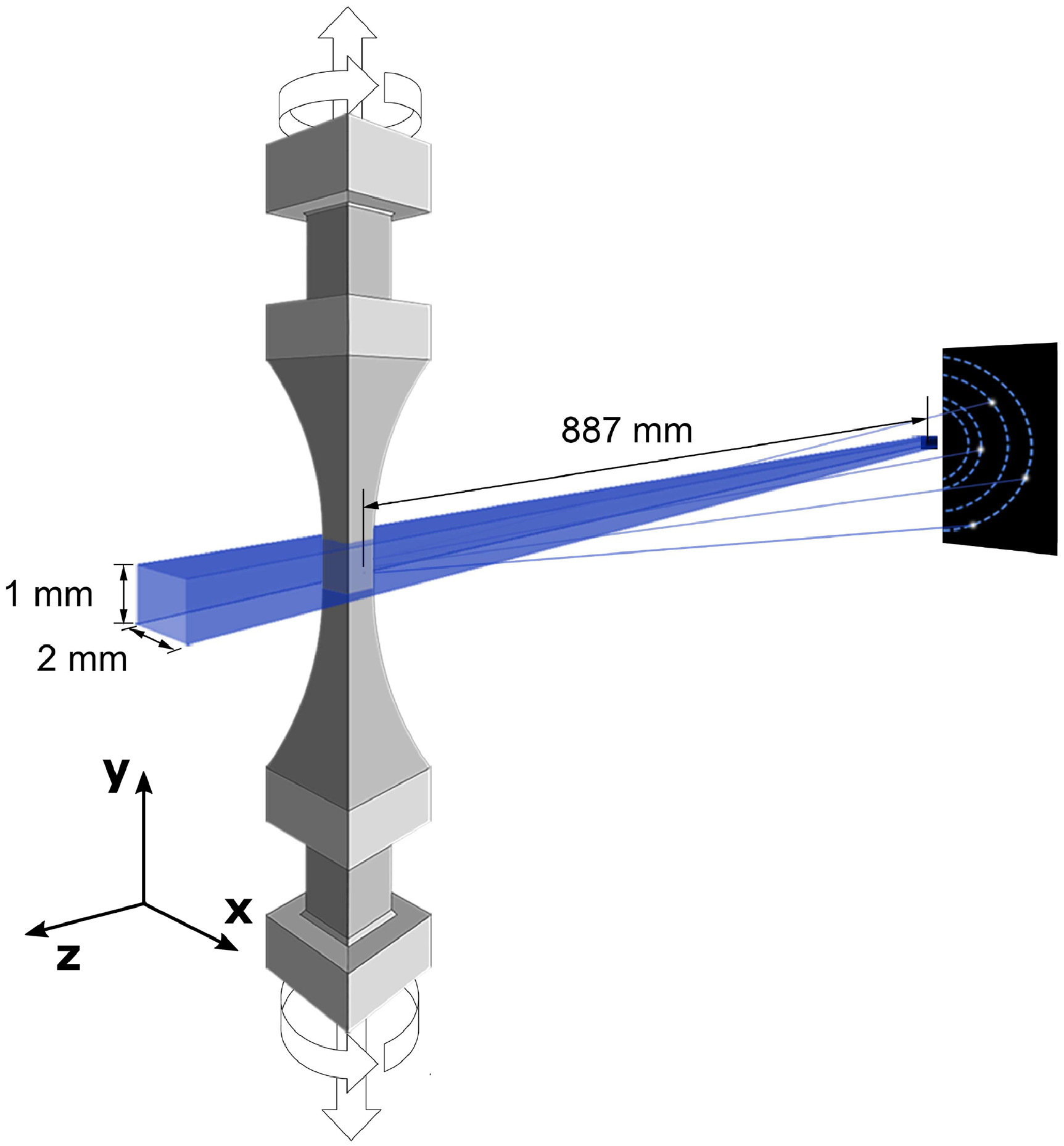 The study of plasticity in engineering alloys is primarily a study of dislocation motion. For data analysis, the unsupervised learning technique is applied to the analysis of X-ray diffraction data measured in-situ during the uniaxial plastic deformation of an additively manufactured nickel-based superalloy. With the aid of a physics-based material model, the authors find that defect densities dictate strength and plastic flow behavior in the alloy.
The study of plasticity in engineering alloys is primarily a study of dislocation motion. For data analysis, the unsupervised learning technique is applied to the analysis of X-ray diffraction data measured in-situ during the uniaxial plastic deformation of an additively manufactured nickel-based superalloy. With the aid of a physics-based material model, the authors find that defect densities dictate strength and plastic flow behavior in the alloy.
Acta Materialia 181, 510 (2019); DOI: 10.1016/j.actamat.2019.10.011
Crystallization-Driven Self-Assembly of Coil−Comb-Shaped Polypeptoid Block Copolymers
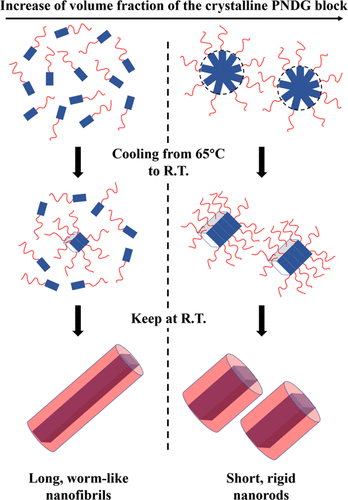 Crystallization-driven self-assembly polymers into well-defined nanoscopic structures with different morphologies and functionalities has attracted increasing attention. Using X-ray solution scattering in combination with other techniques it is possible to understand the relationship between chemical composition, micellar morphology, and crystallization-driven self- of coil−comb-shaped diblock copolypeptoids which is an important step toward the rational design of anisotropic polymeric nanostructures with tailorable morphology.
Crystallization-driven self-assembly polymers into well-defined nanoscopic structures with different morphologies and functionalities has attracted increasing attention. Using X-ray solution scattering in combination with other techniques it is possible to understand the relationship between chemical composition, micellar morphology, and crystallization-driven self- of coil−comb-shaped diblock copolypeptoids which is an important step toward the rational design of anisotropic polymeric nanostructures with tailorable morphology.
Macromolecules 52, 8867 (2019); DOI: 10.1021/acs.macromol.9b01546