What did the Scientists Discover?
Serial protein microcrystallography (SMX) is a promising technique to determine the structure of biological macromolecules, especially in cases where large crystals are difficult to obtain. In the past, SMX has typically been performed at x-ray free electron lasers (xfels). Unfortunately, xfel time is rare and expensive; hence, there is great interest in doing SMX at more readily available x-ray storage ring sources (SRS). A major limitation of SMX at SRS is that radiation damage limits the minimum size of the crystals that may be used.
In 2017 Martin-Garcia et al. (IUCrJ, 4 (2017) 439) solved the structure of a protein using microcrystal data acquired at the Advanced Photon Source, a SRS near Chicago. Most of the crystals were too small to provide interpretable data and the data from these crystals were discarded as too noisy to be analyzed. Instead, the structure was determined from the very small fraction of larger crystals in the SMX experiment.
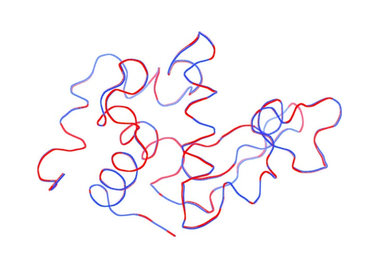
Cornell University and CHESS scientists have developed a method to analyze x-ray SMX data that have previously been thought to be too weak to be of use. In this paper, the Cornell method was applied to the data that Martin-Garcia et al. had discarded as unusable. The resultant structure was comparable to that determined from the larger crystals.
Impact:
The experiment has shown that SMX can be successfully performed at SRS using much smaller crystals than formerly thought possible, thereby greatly expanding the number of protein structures that can be determined using SRS that are available world-wide. This will advance the pace of biomedical discovery.
Collaborators:
Sol M. Gruner, Laboratory of Atomic and Solid State Physics, Physics Department, Cornell University, email: smg26@cornell.edu
Ti-Yen Lan, Laboratory of Atomic and Solid State Physics, Physics Department, Cornell University,
Jennifer L. Wierman, Cornell High Energy Synchrotron Source & MacCHESS, Cornell University
Mark W. Tate, Laboratory of Atomic and Solid State Physics, Physics Department, Cornell University,
Hugh T. Phillipp, Laboratory of Atomic and Solid State Physics, Physics Department, Cornell University,
Jose M. Martin-Garcia, School of Molecular Sciences and Biodesign Center for Applied Structural Discovery, Biodesign Institute, Arizona State University
Lan Zhu, School of Molecular Sciences and Biodesign Center for Applied Structural Discovery, Biodesign Institute, Arizona State University
David Kissick, Advanced Photon Source, Argonne National Lab
Petra Fromme, School of Molecular Sciences and Biodesign Center for Applied Structural Discovery, Biodesign Institute, Arizona State University
Robert F. Fischetti, Advanced Photon Source, Argonne National Lab
Wei Liu, School of Molecular Sciences and Biodesign Center for Applied Structural Discovery, Biodesign Institute, Arizona State University
Veit Elser, Laboratory of Atomic and Solid State Physics, Physics Department, Cornell University
Publication citation:
Lan TY, Wierman JL, Tate MW, Philipp HT, Martin-Garcia JM, Zhu L, Kissick D, Fromme P, Fischetti RF, Liu W, Elser V, Gruner SM, "Solving protein structure from sparse serial microcrystal diffraction data at a storage-ring synchrotron source." IUCrJ. 2018 Jul 20;5(Pt 5):548-558. doi: 10.1107/S205225251800903X . eCollection 2018 Sep 1.
Funding:
| Funding Agency | Grant Number |
|---|---|
| US Department of Energy (DOE) |
DE-SC0005827 |
| Taiwan Government | |
| National Science Foundation |
DMR-1332208 |
| National Institute of General Medical Sciences |
GM-103485 |
| DOE |
DE-SC0017631 |
| Center for Applied Structural Discovery (CASD) at the Biodesign Institute at Arizona State University | |
| Flinn Foundation Seed Grant |
1991 |
| STC Program of the National Science Foundation through BioXFEL |
1231306 |
| NIH |
R21 DA042298 R01 GM124152 |
| NSF STC |
1231306 |
| Flinn Foundation Seed Grant | |
| DOE Office of Science |
DE-AC02-06CH11357 |