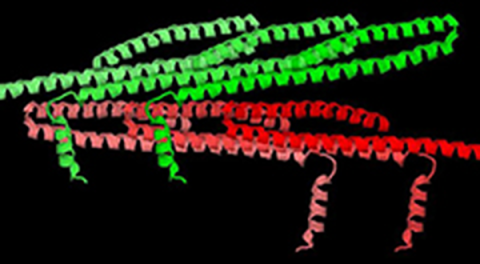
Usually, HPC has negligible effect on the crystal structure. Occasionally, it causes a small change in packing of the molecules in the crystal. For crystals containing a lot of solvent, pressure may cause them to collapse, destroying their diffraction. HPC on crystals of Snf7, however, results in a large change in molecular packing without destroying the crystallinity. In fact, the pressure-cooled crystals diffract better than the normally cooled crystals, in spite of a 30% decrease in unit cell volume!
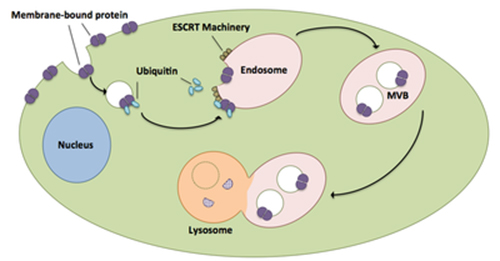
Snf7 is a component of ESCRT-III, which is one of the endosomal sorting complexes required for transport (ESCRTs). These complexes are membrane remodeling machineries that mediate diverse fundamental cellular processes, including the biogenesis of multivesicular bodies (MVBs) during receptor down-regulation, enveloped virus budding, cytokinesis, plasma membrane repair, nuclear pore complex assembly and nuclear envelope reformation. The job of ESCRTs is to package transmembrane proteins tagged with ubiquitin into vesicles that merge with the “late endosome”, creating an MVB that ultimately delivers cargos to the lysosome for destruction (Figure 1). Upstream ESCRT components (ESCRT-0, I & II) assemble into complexes which sort ubiquitinated cargo on the endosomal surface. In addition, ESCRT-II initiates the assembly of the ESCRT-III complex, which together with Vps4 is responsible for remodeling endosomal membranes.
ESCRT-III is a dynamic hetero-polymer, composed of four core subunits (Vps20, Snf7/Vps32, Vps24 & Vps2) and four accessory subunits (Ist1, Did2/Vps46, Vps60 & Chm7). Although all ESCRT-III subunits share a common domain organization, each subunit appears to contribute a specific function. All ESCRT-III subunits are inactive monomers in the cytoplasm; when activated they assemble into spiraling polymers on endosomes to drive cargo sequestration, membrane invagination and constriction.
The group of scientists led by Prof. Scott Emr of Cornell University were able to crystallize and solve the structure of the core region (residues 12-150) of the most abundant ESCRT-III subunit, Snf7 (Snf7core), in its activated conformation. X-ray diffraction data were collected at the F1 beamline at CHESS. At ambient pressure Snf7core crystals diffracted to 2.4 Å at best; 2.7 Å was typical. HPC was used in an attempt to improve this resolution limit. A crystal cryocooled under the typical HPC pressure of 200 Mpa for 30 minutes diffracted out to 1.55Å. Surprisingly, this crystal showed a reduction of about 30% in unit cell volume, due to repacking of the layers of Snf7core molecules (Figure 2). Further HPC experiments on more Snf7corecrystals confirmed the increase in diffraction resolution of pressure-cooled Snf7core crystals (most diffracted to better than 2Å), coupled with a large change in molecular packing. The unit cell dimensions varied as a function of pressure applied in the HPC process.
Unlike crystal structures of other ESCRT-III subunits (Vps24 & Ist1), which have been determined in their inactive conformations and adopt a canonical four α-helical core domain fold (closed conformation) (Figure 3A), the Snf7core contains only three α-helices packed into a highly elongated structure (open conformation) (Figure 3B). The α3 and α4 helices undergo large-scale structural rearrangements from the proposed autoinhibited ESCRT-III fold observed in the structures of Vps24 and Ist1. The open form contains an ~90Å long α-helix combining the α2 and α3 segments that were distinct α-helices in structures of Vps24 and Ist1. The orientation of the α4 helix varies substantially in the structures processed at different pressures, but the open conformation is maintained. The closed-to-open conformational change not only extends a cationic membrane-binding surface, but also exposes hydrophobic and electrostatic protein interacting surfaces for polymerization. This is the first crystal structure of an ESCRT-III subunit in its activated conformation. Using in vitro reconstitution and pulsed dipolar electron spin resonance spectroscopy techniques, the Emr group found that full-length Snf7 adopted the same active (open) conformation, and assembled into ~30Å periodic protofilaments, in a near-native lipid environment.
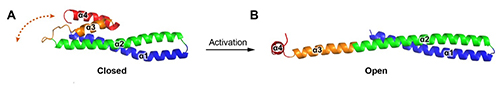
Figure 3: Crystallographically determined structures of (A)Vps24 and (B)Snf7core.
Reference:
S. Tang, W.M. Henne, P.P. Borbat, N.J. Buchkovich, J.H. Freed, Y. Mao, J.C. Fromme & S.D. Emr, “Structural basis for activation, assembly and membrane binding of ESCRT-III Snf7 filaments”, eLife 2015;10.7554/eLife.12548.