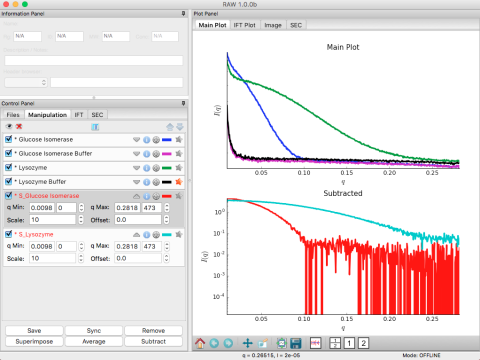
Proteins in solution often stick together in clumps. This can be a serious problem for BioSAXS data because large objects diffract much more strongly at small angles than small objects. When you have large oligomers in solution along with monomeric protein, your Guinier plot will not be linear: no matter how small you take your q range, you won't ever be able to get enough points with q_min*Rg<1.3.
In general, when you see aggregation, you will need to diagnose the problem: can the aggregates be removed by aggressive centrifuging? (rare); do you need to do a fresh size-exclusion chromatography run? Filtration? You might also achieve better results by changing buffer conditions or pH.
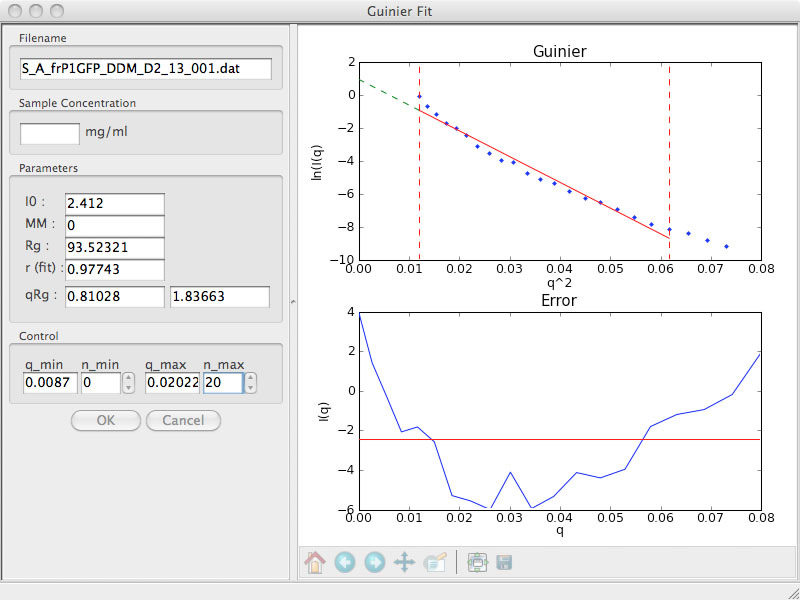
In this plot, notice the characteristic "smile" shape in the Error plot. Unfortunately, this is unusable data. But these users were lucky. They centrifuged their sample at high speed (70,000 rpm in this case) for 30 min and look what happened to the Guinier plot:
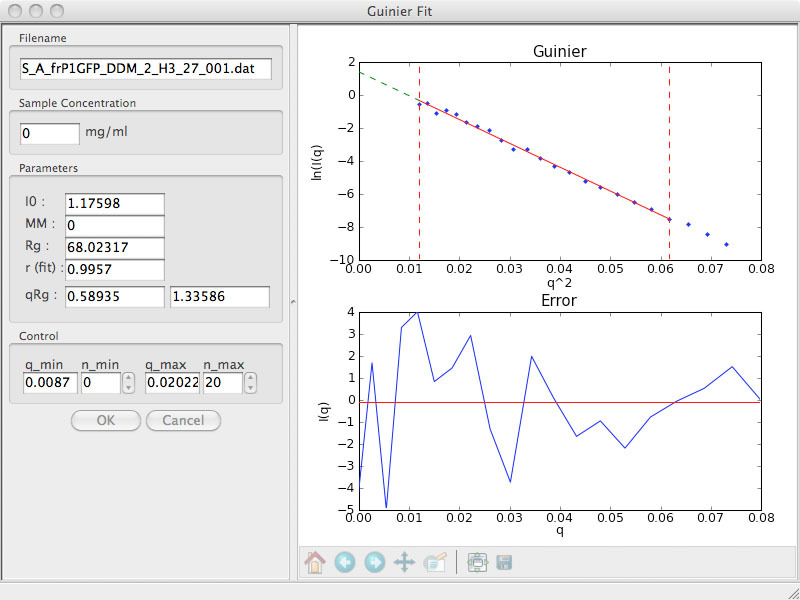
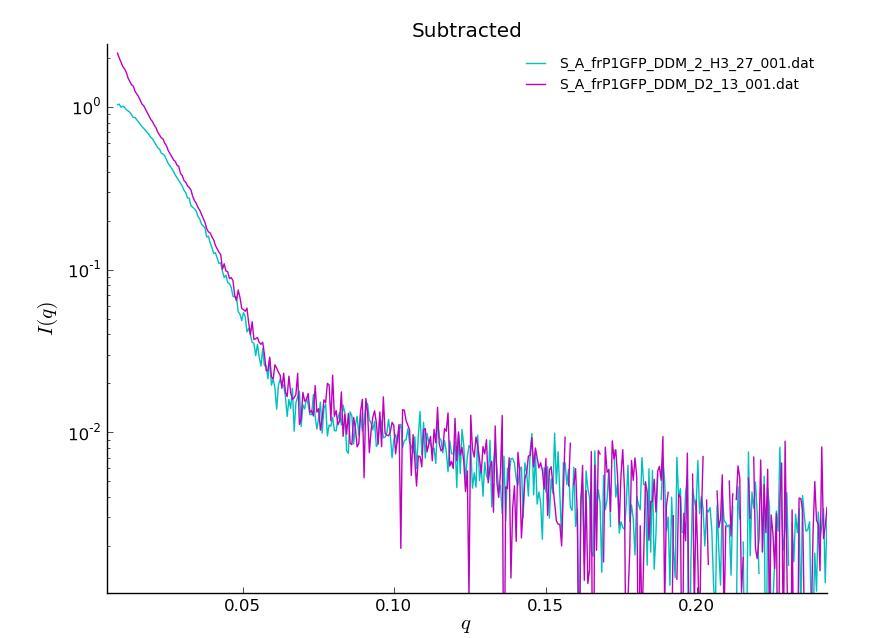
Now, the curve is beautifully linear and there are 20 points with q*Rg<1.33. You can see the dramatic change when you compare scattering curves before and after centrifuging: